Identifier types are present in the database:Ī search is also performed on the sequence descriptor line. Type a gene name, GI number, Uniprot code, free text etc. Current Phyre2 server load = 0% (normal running) Will give you the answer you require in a fraction of the time. For most users, most of the time, 'normal mode' Please do not use "intensive mode" unless your search using "normal mode" indicates that a single model does not cover most of your sequence. Use "batch" processing (under Expert Mode). If you have more than 5 or 6 sequences to model, it is easier for you (and better for everyone!) if you Sequence against your own in-house structures and those from the Protein Data Bank, as well as models downloaded from the AlphaFold Protein Structure Database. 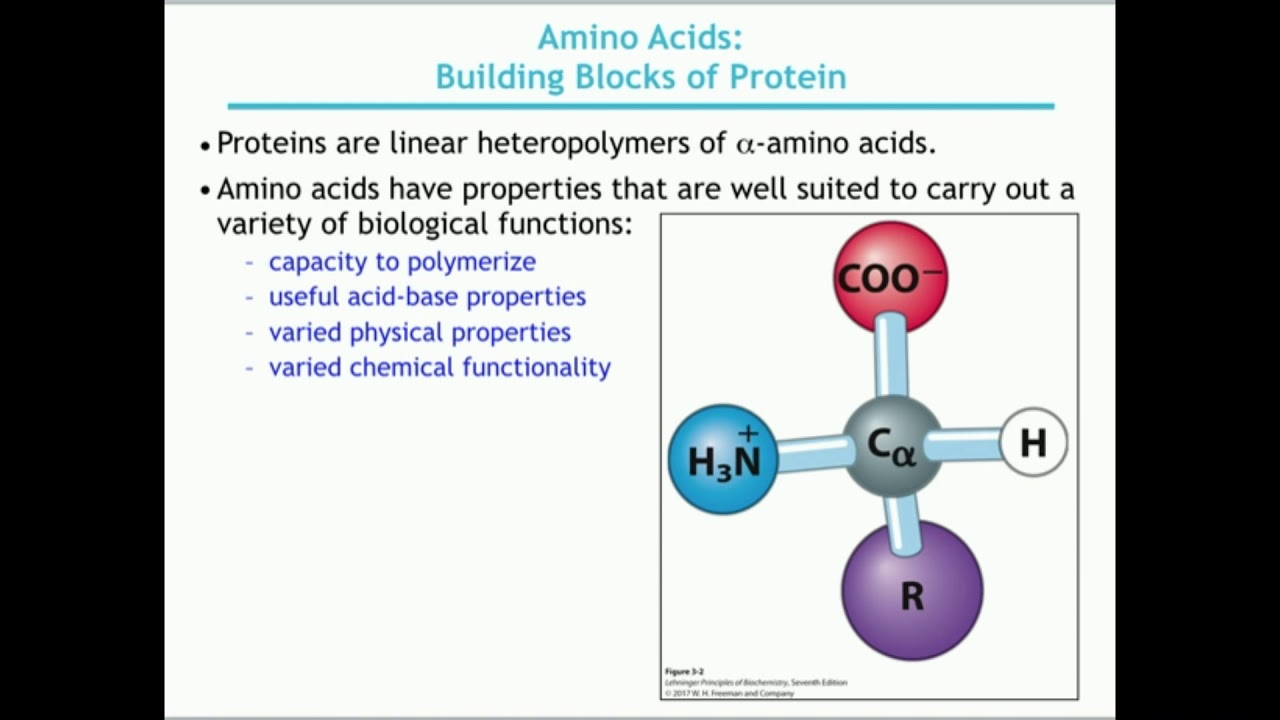
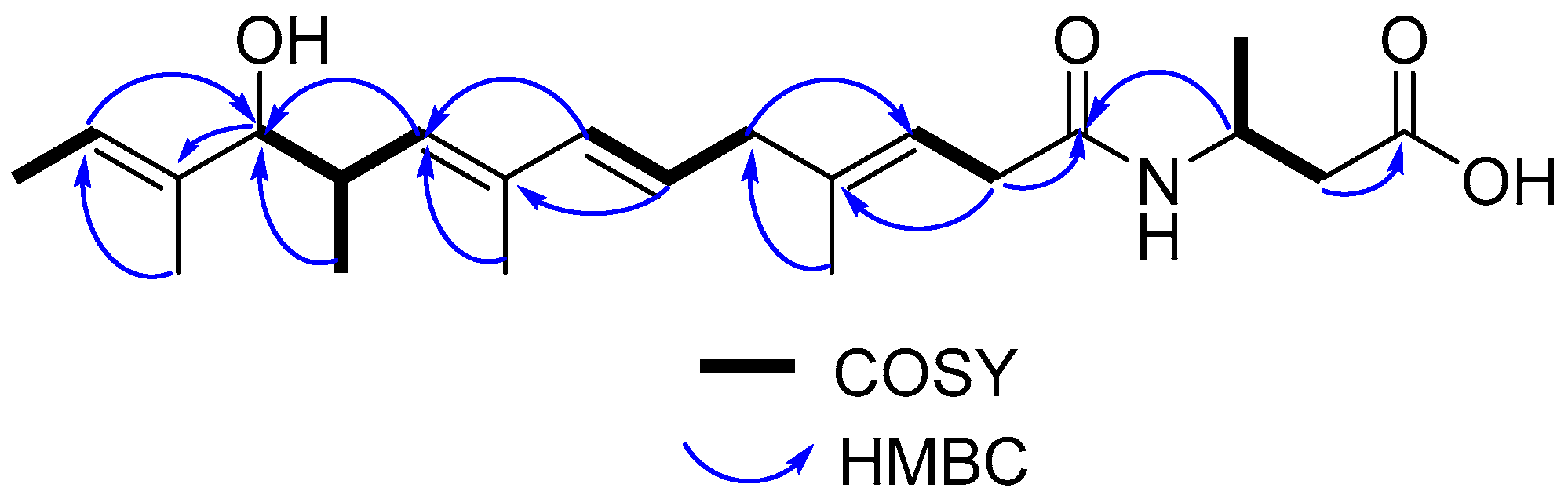
You can use "One-to-One Threading" to model your Registering for this is quick and easy on our Login page. Log in (top left-hand corner of this web page) to use our Expert Mode features, including batch job submission and One-to-One threading. H C N = O C = O Alpha helix Every C=O bonded to N-H 4 residues away forms a helix core is backbone R-groups outside 3.Follow Homology/analog Y Recognition Engine V 2.0 H H H _ _ _ _ H N1- C1- C1- N2- C2- C2- N3- C3- C3- OH = O O O R R R _ _ _ Secondary structure hydrogen bonding backbone groups H-bond donors H-bond acceptors -helix Two main secondary structures: -sheet Phe-Val-Asn-Gln-His Gln-His-Leu-Cys His-Leu-Cys-Gly-Ser His-Leu-Val-Glu Gly-Ser-His-Leu-Val Leu-Val-Glu-Ala Phe-Val-Asn-Gln-His Gln-His-Leu-Cys Leu-Val-Glu-Ala His-Leu-Cys-Gly-Ser Gly-Ser-His-Leu-Val His-Leu-Val-Glu Primary structure study evolution -chain 146 residues horses - humans = 26 pigs - humans = 10 gorillas - humans = 1 1 successful change / 10,000,000 years Primary structure - selective hydrolysis 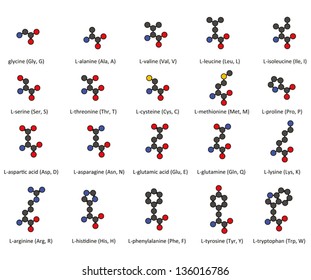
Primary structure sequence of amino acids hemoglobin transports O2 and CO2 300 amino acids 4 protein chains Sickle cell anemia 6thamino acid from N-terminus Glu R Val -CH2CH2-CO2H -CH(CH3)2 water soluble water insoluble H H H _ _ _ = O O O R R R _ _ _ Polypeptides “backbone” _ H N1- C1- C1- N2- C2- C2- N3- C3- C3- OH peptide bonds C-terminal residue N-terminal residue biological activity = structure 4 levels protein structure Amino acids R-groups non-polar polar acidic basic proteins condensation between carboxylic acids and amines + + H2O carboxylic acid amide amineĪmides resonance structure amides dipeptide glycine alanine Ala-Gly +H2O 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
